IVILAB Research Projects
:: CompTIES :: PSI :: objects :: rooms :: plants :: neurons :: fungus :: vision_and_language :: trails :: SLIC :: tracking ::
See also: IVILAB Undergraduate research
Computational Temporal Emotion Systems
This research brings theory-driven Bayesian modeling and inference into the domain of temporal emotional interactions within personal relationships. We are also developing a shared computational infrastructure for researchers in this domain in collaboration with iPlant. The iPlant project is an NFS funded computational infrastructure project initially focused on plant biology. For more details on compTIES, please see the project website . CompTIES is in collaboration between Emily Butler (PI, Family Studies and Human Development), Kobus Barnard (IVILAB / CS, Clay Morrison (IVILAB / SISTA), Matthias Mehl (Psychology).
![]() Funded by NSF grant BCS-1322940.
Funded by NSF grant BCS-1322940.
Link to: 1) Project website and 2) Our NSF funded workshop.
Understanding activities from video: Persistent Stare through Imagination (PSI)
 Inferring human activity from images using conventional approaches (e.g.,
classification based on low level features) is difficult because meaningful
activity concepts are abstract at the semantic level and exhibit a very wide
range of visual characteristics. Consider the verb "follow." A person may follow
another person, a car may follow a motorcycle, a dog may follow its owner, so
on. Further, whether the activity "follow" is actually better described as
"chase" includes intent which is also not directly linked to low level features.
In this project the goal is to develop methods for deep understanding of
activity from video.
Inferring human activity from images using conventional approaches (e.g.,
classification based on low level features) is difficult because meaningful
activity concepts are abstract at the semantic level and exhibit a very wide
range of visual characteristics. Consider the verb "follow." A person may follow
another person, a car may follow a motorcycle, a dog may follow its owner, so
on. Further, whether the activity "follow" is actually better described as
"chase" includes intent which is also not directly linked to low level features.
In this project the goal is to develop methods for deep understanding of
activity from video.
The image shows an alignment of an activity video (person walking) and a physics based simulator (Cartwheel3D). If we can understand the movie in terms of physics, then we can make natural inferences such as a box that is kicked out the way is light, and one that is tripped over is probably heavy.
This project is a collaboration between researchers specializing at different levels of representation: 1) low level feature based recognition (Deva Ramanan's group at UCI), 2) mid level representations of what actors are in the scene and their trajectories (IVILAB), 3) learning behaviour models from image data (the SISTA robotics lab led by Ian Fasel), and 4) language level semantic understanding of activities (the SISTA AI lab led by project PI, Paul Cohen).
Link to: 1) ICCV 2013 paper.
Funded by DARPA through the Mind's Eye Program
Learning Models of Object Structure
This project is developing approaches for learning stochastic 3D geometric models for object categories from image data. Representing objects and their statistical variation in 3D removes the confounds of the imaging process, and is more suitable for understanding the relation of form and function and how the object integrates into scenes.
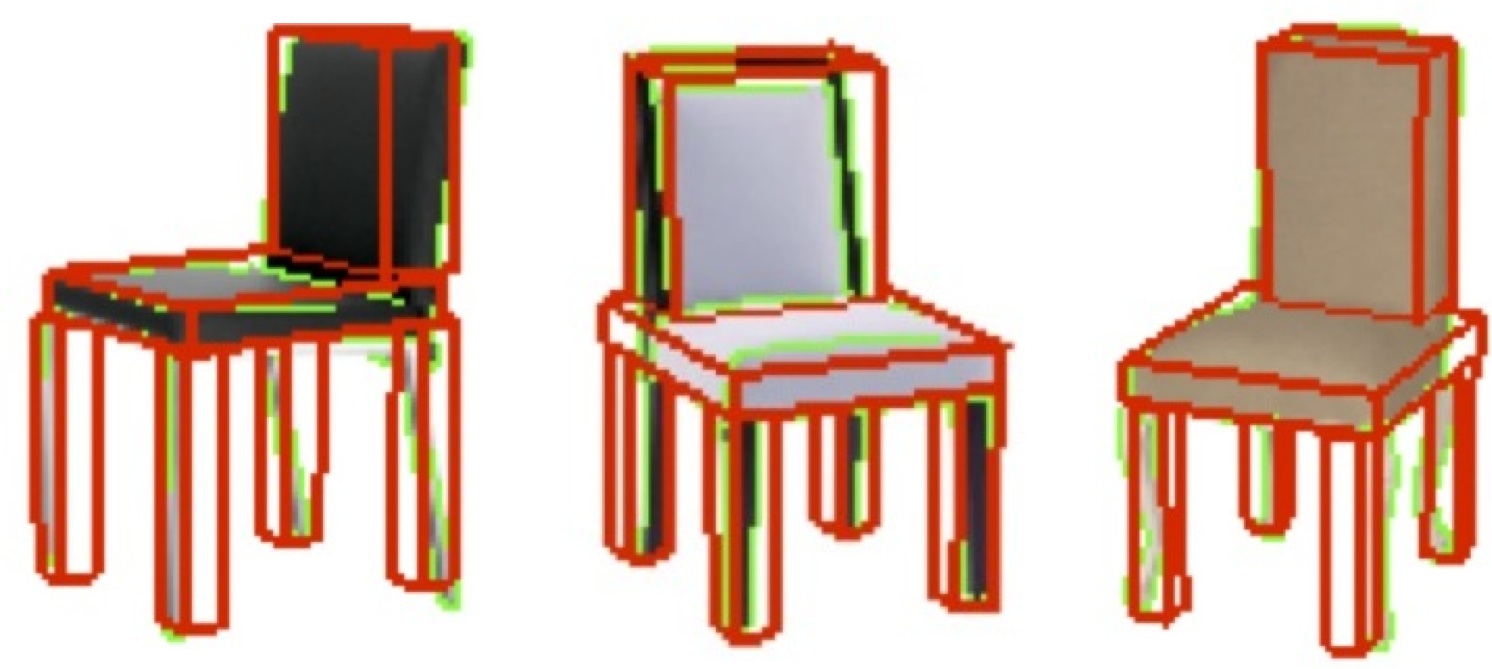 The image to the right shows a simple model for chairs learned from a modest set
of 2D images using the representation of a collection of connected blocks and
the key assumption that the topology is consistent across the
object category. The particular instances that are fit collaterally are shown in
red. For the category we learn the topology and the statistics of the block
parameters.
The image to the right shows a simple model for chairs learned from a modest set
of 2D images using the representation of a collection of connected blocks and
the key assumption that the topology is consistent across the
object category. The particular instances that are fit collaterally are shown in
red. For the category we learn the topology and the statistics of the block
parameters.
Initial work formed the bulk of Joseph Schlecht's dissertation.
Link to: 1) Project page, and 2) NIPS'09 paper.
![]() Funded by NSF CAREER grant IIS-0747511.
Funded by NSF CAREER grant IIS-0747511.
Generative Modeling of Indoor Scenes
 In conjunction with the previous discussed project, we would like to understand
scenes in geometric and semantic terms---what is where in 3D. Doing this from a
single 2D image involved inferring the parameters of the camera, which can be
done assuming a strong model. In this case we adopt the Manhattan world
assumption, namely that most long edges are parallel to three principle axes.
Different from other work, we develop a generative statistical model for scenes,
and rely only on detected edges for inference (for now).
In conjunction with the previous discussed project, we would like to understand
scenes in geometric and semantic terms---what is where in 3D. Doing this from a
single 2D image involved inferring the parameters of the camera, which can be
done assuming a strong model. In this case we adopt the Manhattan world
assumption, namely that most long edges are parallel to three principle axes.
Different from other work, we develop a generative statistical model for scenes,
and rely only on detected edges for inference (for now).
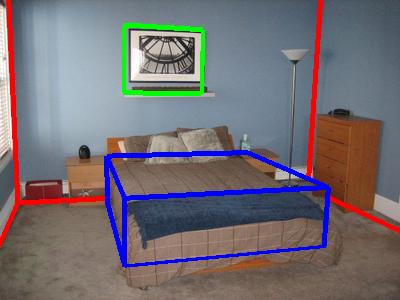

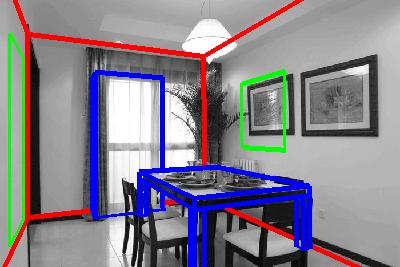
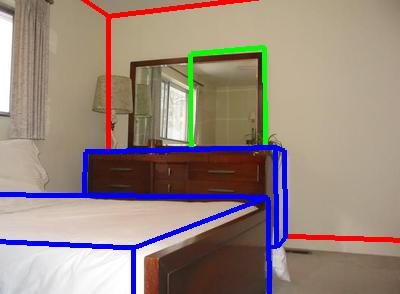
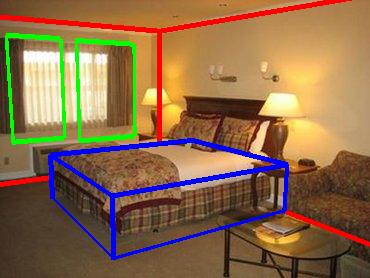 The image shows fits for two rooms. Red boxes are room boundaries, green boxes
are frames (pictures, windows, doors), and the blue boxes are furniture bounding
boxes.
The image shows fits for two rooms. Red boxes are room boundaries, green boxes
are frames (pictures, windows, doors), and the blue boxes are furniture bounding
boxes.
Contributions to this work have been made by Luca del Pero, Joseph Schlecht, Ernesto Brau, Jinyan Guan, and undergraduates Emily Hartley, Bonnie Kermgard, Joshua Bowdish, Daniel Fried, and Andrew Emmott.
Link to: CVPR'13 paper, and data.
Link to: CVPR'12 paper, and data
Link to: CVPR'11 paper, and data.
![]() Funded by NSF CAREER grant IIS-0747511.
Funded by NSF CAREER grant IIS-0747511.
Plant Structure from Images
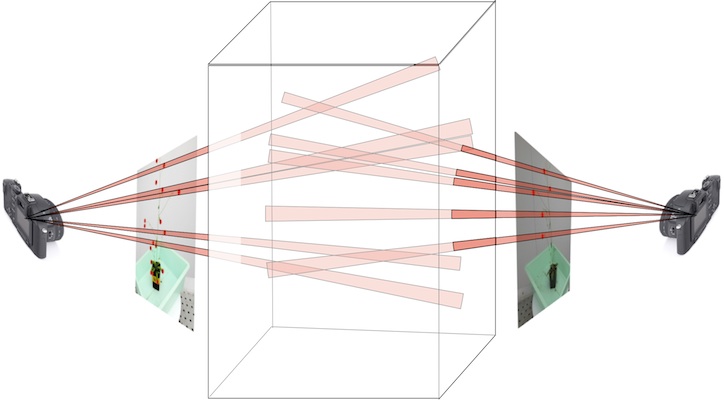
Quantifying plant geometry is critical for understanding how subtle details in form are caused by molecular and environmental changes. Developing automated methods for determining plant structure from images is motivated by the difficulty of extracting these details by human inspection, together with the need for high throughput experiments where we can test against a large number of variables.
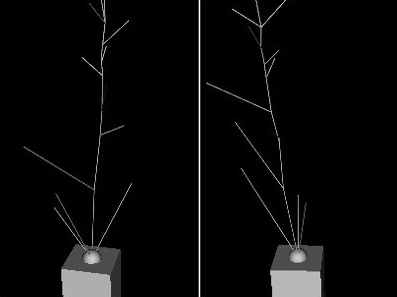
To get numbers for structure we fit geometric models of plants to image data. The top picture shows how features detected in multiple views from calibrated cameras can provide hypothesis for structure that become "data-driven" samples in an MCMC approach to fitting structure. The next row shows two views (left), and corresponding fits of the skeleton to the image data, projected using camera models corresponding to those two views. The heavy lifting on this project is being done by PHD student Kyle Simek.
 This project is in collaboration with the
Palanivelu Lab
and
Rod Wing's group.
This project is in collaboration with the
Palanivelu Lab
and
Rod Wing's group.
Funding.
Kyle has held
a Department of Education GAANN fellowship during the first part of this project. He has been helped by undergraduates
funded by
the NSF
iPlant
project via
Morphology of Brain Neurons
 We are developing models for the structure of Drosophila brain neurons grown in
culture. Genetic mutations and/or environmental factors can have a big impact on
neuron morphology, which in turn effects their function. Automated methods for
quantifying the neuron structure will enable large scale screens for drugs that
compensate for genetic defects or mitigate the effects of toxins.
We are developing models for the structure of Drosophila brain neurons grown in
culture. Genetic mutations and/or environmental factors can have a big impact on
neuron morphology, which in turn effects their function. Automated methods for
quantifying the neuron structure will enable large scale screens for drugs that
compensate for genetic defects or mitigate the effects of toxins.
The image (*) shows an example of a wild type neuron (left), and a "filagree" phenotype associated with a particular mutation (right). Notice that the filagree neurites exhibit significantly more curvature. We are able to distinguish individuals from these populations relatively reliably using image processing methods to quantify the curvature. However, due to the huge variation in phenotype, developing such a method for each one is impractical. Rather, we are pursuing a structural modeling approach, together with statistical inference to learn the model parameters for each phenotype, and to fit learned models to individuals not used for learning.
(*) Image source: Kraft et al., J. Neuroscience, 2006.
This work in collaboration with Linda Restifo and her research group.
Fungal Structure from Image Stacks

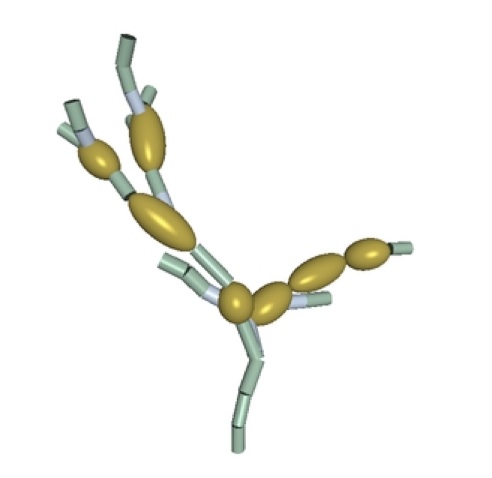 Our goal in this project is to learn and fit structure models for microscopic
filamentous fungus such as those from the genus Alternaria. The data are image
sequences taken at stepped focal depths (stacks). The left hand image pair shows
two images out of 100. Notice the significant blur due to the limited depth of
field. We deal with the blur and exploit the information hidden in it by
fitting the microscope point spread function together with the model. The right
hand images shows the fit model with the cylinders corresponding to hyphae, and
the ellipsoids representing spores.
Our goal in this project is to learn and fit structure models for microscopic
filamentous fungus such as those from the genus Alternaria. The data are image
sequences taken at stepped focal depths (stacks). The left hand image pair shows
two images out of 100. Notice the significant blur due to the limited depth of
field. We deal with the blur and exploit the information hidden in it by
fitting the microscope point spread function together with the model. The right
hand images shows the fit model with the cylinders corresponding to hyphae, and
the ellipsoids representing spores.
Initial heavy lifting was done by PHD student Joseph Schlecht, guided by modeling work by undergraduate Kate Taralova (see below). Undergraduate Johnson Truong has worked on a fast approach for computing the blurring for model hypotheses.
Link to: 1) CVPR'07 paper.
This project is in collaboration with the Pryor lab.
Linking Vision and Language
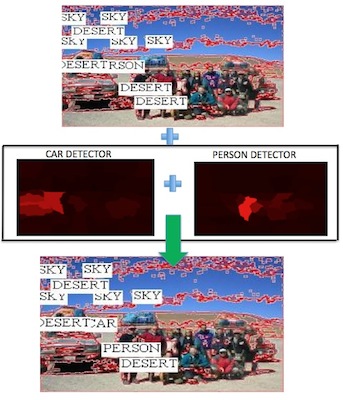
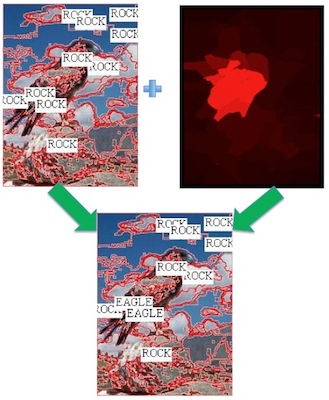 Image keywords and captions provide information about what is in the image, but
we do not know which words correspond to which image elements. Alternatively,
words may provide complementary non-visual information. Dealing with this
correspondence problem enables using such data as training data
for image understanding and object recognition, and can improve multi-modal
searching, browsing, and data mining.
Image keywords and captions provide information about what is in the image, but
we do not know which words correspond to which image elements. Alternatively,
words may provide complementary non-visual information. Dealing with this
correspondence problem enables using such data as training data
for image understanding and object recognition, and can improve multi-modal
searching, browsing, and data mining.
Recent IVILAB work on this topic has focused on aligning image caption or key words with visual features. The images on the right show how specialized object detectors can help improve this (labels for only the 10 largest regions are shown). The second example (right) shows that the object detector for "bird" can be used to align "eagle", given that we can look up the relation between the two using WordNet. Heavy lifting is being done by PHD student Luca del Pero, with a lot of help from undergraduate students Philip Lee, Emily Hartley, and James Magahern.
Link to: 1) ACM MM'11 paper, 2) CVPR'07 paper.
Link to our "words and pictures" research since 2000.
![]() Support for this project has come from NSF, ONR, and TRIFF.
Support for this project has come from NSF, ONR, and TRIFF.
Finding Trails in Satellite Photos
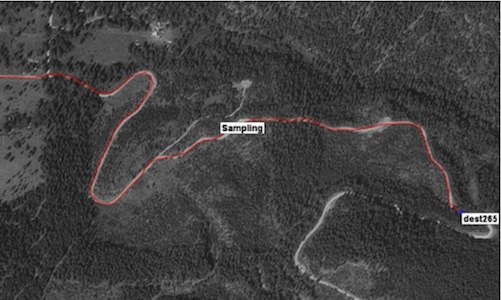 The image to the right shows a satellite photo of a trail with an automatically
extracted trail overlaid in red (end points are provided). In this image the
method works well, but in general, this is a difficult task as trails are often
faint, obscured by trees, and similar in appearance to dry stream beds. Trails
both deteriorate over time and also appear due to human activity. The volatile
nature of trails makes extracting them from satellite photographs important. In
this work we exploit GPS tracks of travellers on trails to train and validate
automated methods for extracting them. This work began with
Scott Morris's
PHD work, and is being extended by
Andrew Predoehl.
The image to the right shows a satellite photo of a trail with an automatically
extracted trail overlaid in red (end points are provided). In this image the
method works well, but in general, this is a difficult task as trails are often
faint, obscured by trees, and similar in appearance to dry stream beds. Trails
both deteriorate over time and also appear due to human activity. The volatile
nature of trails makes extracting them from satellite photographs important. In
this work we exploit GPS tracks of travellers on trails to train and validate
automated methods for extracting them. This work began with
Scott Morris's
PHD work, and is being extended by
Andrew Predoehl.
Link to: 1) CVPR'13 paper. 2) CVPR'08 paper.
Funding. Both Scott and Andrew have held Department of Education GAANN fellowships while working on this project.
Semantically Linked Instructional Content (SLIC)
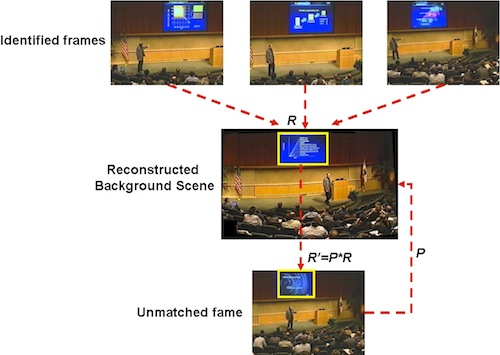 The goal of this project is to make instructional video more accessible to
browsing and searching. A key tool for this is to match slide images from
presentation files (PPT/PPTX, Keynote, PDF) to videos that use them. We approach this
problem by using SIFT keypoint matching under a consistent homography. The
figure shows how we can use initial matches to build a background
model, which in turn helps us match more difficult frames because we now know
where the slide image must be.
The goal of this project is to make instructional video more accessible to
browsing and searching. A key tool for this is to match slide images from
presentation files (PPT/PPTX, Keynote, PDF) to videos that use them. We approach this
problem by using SIFT keypoint matching under a consistent homography. The
figure shows how we can use initial matches to build a background
model, which in turn helps us match more difficult frames because we now know
where the slide image must be.
For much more information on the project, including publications and a live demo, see the SLIC project page.
![]() SLIC is partly supported by NSF grant EF-0735191.
SLIC is partly supported by NSF grant EF-0735191.
Simultaneously tracking many objects with applications to biological growth
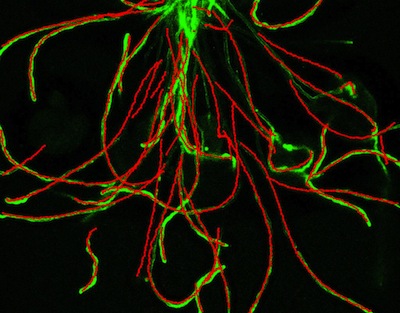 Simultaneously tracking many objects with overlapping trajectories is hard
because you do not know which detections belong to which objects. The vision lab
has developed a new approach to this problem and has applied it to several kinds
of data. For example, the image to the right shows tubes that are growing out of
pollen specs (not visible) towards ovules (out of the picture) in an
Simultaneously tracking many objects with overlapping trajectories is hard
because you do not know which detections belong to which objects. The vision lab
has developed a new approach to this problem and has applied it to several kinds
of data. For example, the image to the right shows tubes that are growing out of
pollen specs (not visible) towards ovules (out of the picture) in an
We are currently working on a statistical model for pollen tube / ovule interaction behaviour given the tracks. This work is being led by PHD student Ernesto Brau with help from undergraduate researcher Phil Lee.
Link to: 1) CVPR'11 paper, 2) Data, and 3) Code.
This project is in collaboration with the Palanivelu lab.
![]() Funding for this project provided by NSF grant IOS-0723421.
Funding for this project provided by NSF grant IOS-0723421.
Identifying machine parts using CAD models
 CAD models provide the 3D structure of many man made objects such as machine
parts. This projects aims to find objects in images based on these models.
However, since the data is most readily available as triangular meshes, 3D
features that are useful for matching 2D images must be extracted from mesh
data. Having done that, we need fast methods to match to them to images.
Applications of this work include identifying parts in machinery maintenance
facilities and finding possible manufactured parts based on user sketches.
CAD models provide the 3D structure of many man made objects such as machine
parts. This projects aims to find objects in images based on these models.
However, since the data is most readily available as triangular meshes, 3D
features that are useful for matching 2D images must be extracted from mesh
data. Having done that, we need fast methods to match to them to images.
Applications of this work include identifying parts in machinery maintenance
facilities and finding possible manufactured parts based on user sketches.
This project is being let by PHD student Luca del Pero, with a lot of help from undergraduates Emily Hartley and Andrew Emmott.
![]() Funding for undergraduates provided by
NSF Grant 0758596 and
REU supplement to NSF
CAREER
grant IIS-0747511.
Funding for undergraduates provided by
NSF Grant 0758596 and
REU supplement to NSF
CAREER
grant IIS-0747511.
Back to top
Inactive and Subsumed Projects
Modeling and visualizing Alternaria
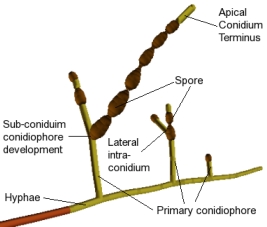 The image to the right is a labeled model of the fungus Alternaria generated by a
stochastic L-system built by undergraduate researcher
Kate Taralova.
For more information, follow this
link. This project continues as part of the plant modeling project.
The image to the right is a labeled model of the fungus Alternaria generated by a
stochastic L-system built by undergraduate researcher
Kate Taralova.
For more information, follow this
link. This project continues as part of the plant modeling project.
This project is in collaboration with the Pryor lab.
![]() Support for undergraduates provided by TRIFF and a REU supplement to a
department of computer science NSF research infrastructure grant
Support for undergraduates provided by TRIFF and a REU supplement to a
department of computer science NSF research infrastructure grant
Evaluation methodology for automated region labeling
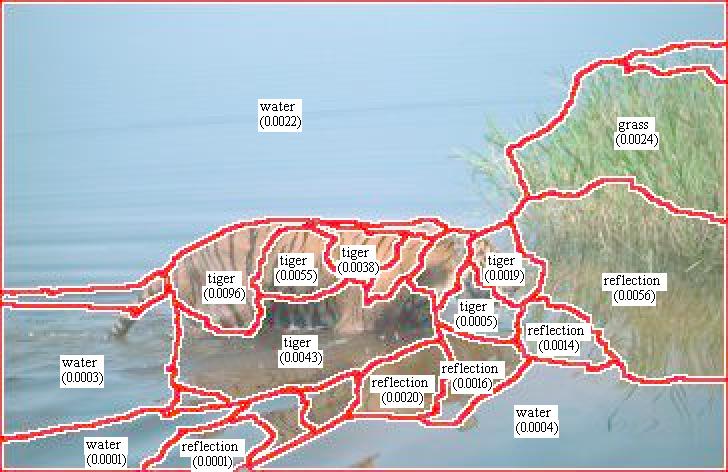 We have developed a scoring system which gives each ground truth region a score
for every word. To apply these maps, we use them to provide scores for any
given, typically imperfect, segmentation used by region labeling algorithms. The
image to right shows a normalized cuts based segmentation of an image together
with the score for the maximal scoring word which is also used to label the
image. Notice that the reflection of the tiger is merged with the tiger, and so
this region will have non-negligible score for both tiger and water, and
necessarily less that than a pure tiger region of comparable size. The base
scoring attached to the ground truth labeling takes into account the frequency
of the concept in the training data and the specificity of the word ("tiger"
scores more than "cat") computed using
WordNet.
This approach allows us to compute both "frequency correct" and "semantic range"
scores on pure visual information, which are honest with respect to prior
probabilities and the trade-off between specificity and expected rate of
mistakes. The "frequency correct" score rewards getting lots of labels correct,
which is often the result of doing well on common words such as "sky" and
"water". The semantic range score provides equal weight for each word, and
provides a measure of how well systems handle rare words.
We have developed a scoring system which gives each ground truth region a score
for every word. To apply these maps, we use them to provide scores for any
given, typically imperfect, segmentation used by region labeling algorithms. The
image to right shows a normalized cuts based segmentation of an image together
with the score for the maximal scoring word which is also used to label the
image. Notice that the reflection of the tiger is merged with the tiger, and so
this region will have non-negligible score for both tiger and water, and
necessarily less that than a pure tiger region of comparable size. The base
scoring attached to the ground truth labeling takes into account the frequency
of the concept in the training data and the specificity of the word ("tiger"
scores more than "cat") computed using
WordNet.
This approach allows us to compute both "frequency correct" and "semantic range"
scores on pure visual information, which are honest with respect to prior
probabilities and the trade-off between specificity and expected rate of
mistakes. The "frequency correct" score rewards getting lots of labels correct,
which is often the result of doing well on common words such as "sky" and
"water". The semantic range score provides equal weight for each word, and
provides a measure of how well systems handle rare words.
Link to: 1) IJCV paper and 2) Data and Code.
Funding provided by TRIFF.
Human based evaluation of content based image retrieval
 Link to:
1)
CVPR'05 paper
and
2)
Data.
Link to:
1)
CVPR'05 paper
and
2)
Data.
Funding provided by TRIFF.
LSST association pipeline
The endeavour to build the Large Synoptic Survey Telescope (LSST) project is a huge undertaking. See lsst.org for details. Not surprisingly, a significant part of the system is the data processing pipeline. We have worked on the temporal linking observations of objects assumed to be asteroids to establish orbits and build data systems for querying the emerging time/space catalog. One of the goals of this research is to help identify potentially hazardous asteroids. This is joint work with Alon Efrat, Bongki Moon, and the many individuals working on the LSST project.Link to: 1) overview paper.
![]() Funding provided by a sub-contract to NSF award #0551161;
Funding provided by a sub-contract to NSF award #0551161;